Editing Plant Genomes
October 29, 2019
Alejandro Hernández, Biotechnology Director for Central America and the Caribbean; CropLife Latin America
Genome editing in plants is achieved thanks to precision biotechnology and it aims to be a milestone in modern agriculture. There are currently applications for permission to sell tomatoes, rice, corn, wheat, soybeans and mushrooms in the United States. Genetic edition promises nutritional changes such as the production of gluten-free wheat or reduced trans fat soybean oils. What is it about?
The genome is the entire genetic material of an organism or a particular species. For higher organisms or eukaryotes, such as plants and animals, there is the nuclear genome and the plastid (plants) and/or mitochondria genome (animals and plant).
The genetic material is presented in highly organized structures in the DNA, and is a genetic code written in four letters: A (adenine) that pairs with T (thymine), and C (cytosine) that pairs with G (guanine).
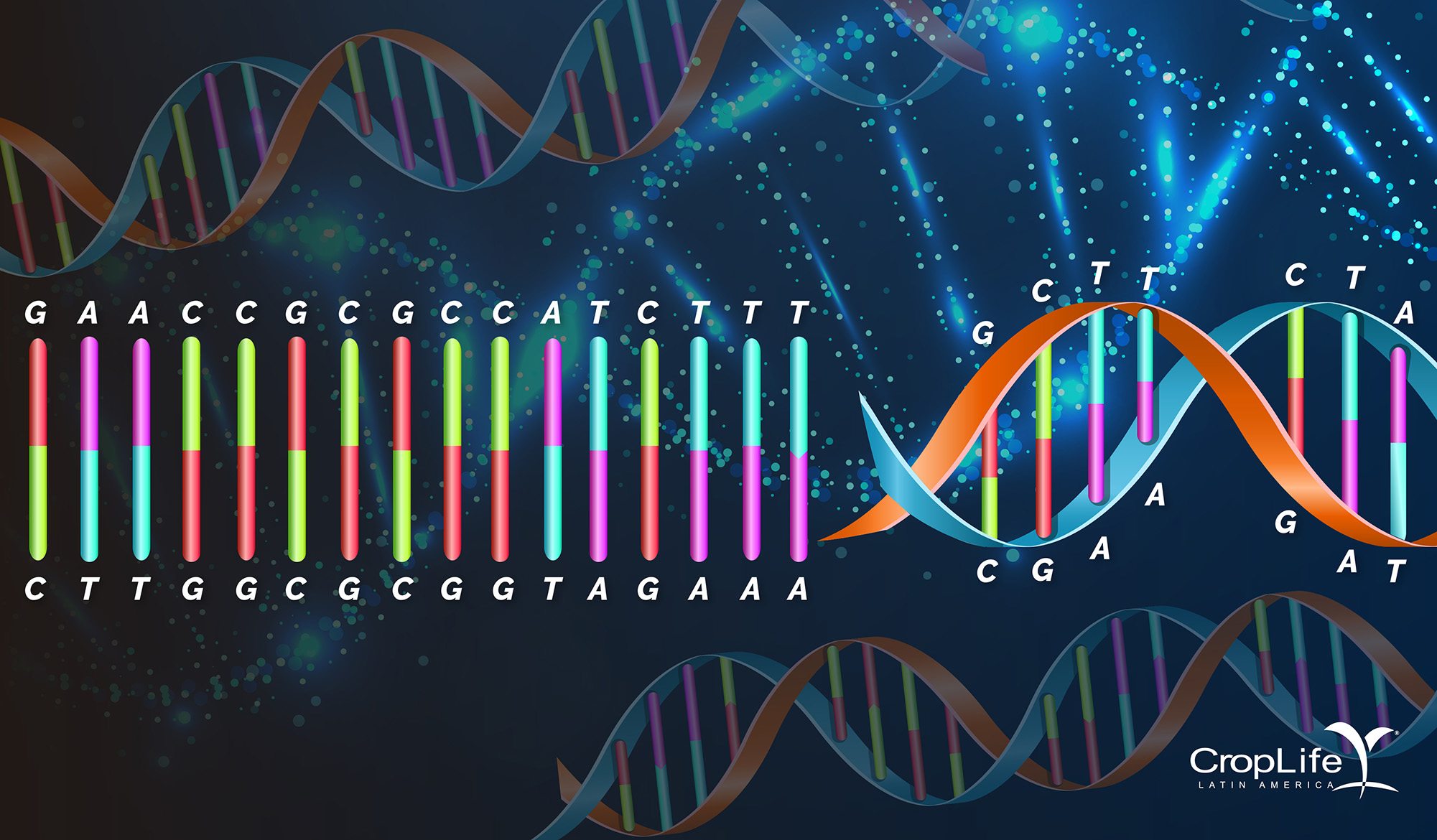

The sequence of a plant genome is similar to a library of letters. Within this library of letters, there is a small proportion that corresponds to genes, or sequences with some function. These genes have small changes that can vary their function. In conventional breeding, these changes are the characteristics that are sought by pollination or random changes. Until only a couple of decades ago we knew very little about genomes and even less about how to adjust, improve or repair them when we found an error. Today, the sequencing tools and new Genome Editing techniques make it possible to generate and verify with extreme precision adjustments in the genetic material. In the case of plants, the scenario is very promising since you can get the same results as with conventional breeding, but in a much lesser time. Therefore, they are called New Breeding Techniques/Technologies (NBTs).
Genome editing technologies are based on 2 principles. The first is that the precise site in the genome must be located in order to perform some function. The most common, but not the only one, is to cut the double strand of DNA ("double-strand breaks" (DSBs)). The second principle is the use of natural genome repair mechanisms. In higher organisms or eukaryotes it is called homologous repair when an identical backup copy is used to repair the genome; or non-homologous when the genome is repaired without the help of a repair strand. The final result is a DNA sequence with repairs or adjustments, which in the literature were originally called “SDN” or “SSN” techniques (“site directed nucleases” or “site specific nucleases” 1, 2 and 3).

Let's explore these two concepts to understand the state of the art of the new genome editing techniques.
Principle 1. Locate the right place in the genome
The first methods to identify some genetic material were found in the 80s as defense mechanisms against viruses. Imagine a pair of DNA scissors, which at that time were called endonucleases and meganucleases, the first had the ability to recognize about 6 to 8 letters and the second 12-40 letters; both could make a cut in the DNA. In the 90s and early 2000, progress was made with the generation of Zinc finger nucleases (recognize 6-18 base pairs) and TALENS (Transcription-Activator-Like Effector Nucleases) that recognize 12-31 bp. From 2012 to date, a series of scissors or molecular enzymes called CAS were identified that are associated with libraries of small fragments of approximately 20 base pairs of virus sequences called CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats). The latter are what are called CRISPR-CAS9 / CAS12a, CAS12b, CAS12e, CAS13a, CAS13b, CAS14.
The story of CRISPR-CAS9 is captivating
The enzyme was discovered by Francisco Mojica in 1993 when he studied the microorganism Haloferax mediterranei and noticed a defense system in bacteria against viruses, consisting of two elements, scissors that he called CAS and a library of information that used the scissors to identify the virus. The latter he called CRISPR in allusion to the series of virus sequences that were very orderly arranged one after the other and separated by an identical DNA that was repeated over and over again, hence the acronym CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats).

The discovery went through many years with a low profile, until 2012, when it was verified that the bacterial system could be used to edit genomic DNA. That is when a scientific competition appears at the highest international level between two universities in the United States: Berkeley University on the west coast under the command of two scientists, Jennifer Doudna and Emmanuelle Charpentier and the Harvard-MIT Broad Institute on the east coast, led by scientist Feng Zhang. The discovery by both groups was based on the fact that CAS9 (the scissors) is perfectly complemented by a sequence that could be custom designed called guide RNA (gRNA), which contains 20 letters that it uses as a reference to locate a particular place in the genome, making it extremely specific.
Principle 2, genome repair
Once a cut in the double strand of DNA is made, the cell's natural repair machinery goes into action. The cellular machinery can repair the genome in different ways, and depending on the type of repair it is called “SDN” or “SSN” (“site directed nucleases” or “site specific nucleases” 1, 2 y 3).
SDN or SSN-1 are the product of non-homologous recombination, that is, a repair that does not use a repair strand and results in insertions, deletions or small changes.
SDS-2 use a DNA strand and therefore the repair is homologous, but they do not insert new DNA, rather repairing the message, for example, they can cause changes at the amino acid level of the encoded protein.
SDN-3 or SSN-3 use a repair strand, by homologous recombination and therefore may or may not result in a new insertion of DNA. This is important to understand since it is possible to insert a complete allele at a specific location in the genome. If this insertion is identical to what is achieved by conventional breeding, the end result is indistinguishable from the conventional techniques.
We are in an exciting time where different technologies converge towards sustainable agriculture. Without a doubt, precision biotechnology will play a key role in producing more and in a more balanced way.
Biography
1. Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., & Charpentier, E. (2012). A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. science, 337(6096), 816-821.















